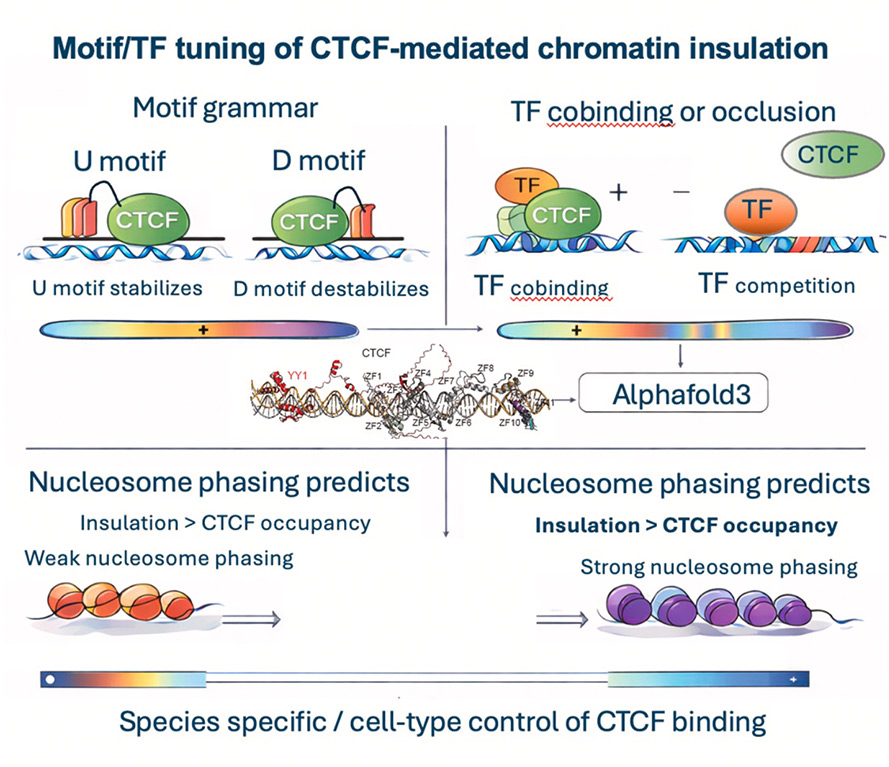
 Motif grammar and transcriptional programs decouple CTCF binding from nucleosome phasing to control cell-type-specific chromatin insulation
Motif grammar and transcriptional programs decouple CTCF binding from nucleosome phasing to control cell-type-specific chromatin insulation
Catherine Do, Guimei Jiang, Paul Zappile, Adriana Heguy, Jane A Skok.
CTCF organizes the genome via cohesin-mediated loop extrusion, insulating topologically associating do- mains (TADs) and constraining enhancer-promoter communication. Yet, CTCF binding varies across cell types and genomic contexts. In mouse, typically accessible sites combine strong motifs with higher CTCF/cohesin occupancy, while inaccessible sites use weaker motifs stabilized by flanking upstream (U) and downstream (D) sequences that engage peripheral zinc fingers to respectively enhance or dampen binding. Notably, TF motifs within ±35 bp bind cooperatively or competitively depending on expression and positional overlap, a finding supported by AlphaFold predictions and allele-specific perturbations. Favorable motif/TF environments can increase CTCF signal yet disrupt nucleosome phasing and weaken in- sulation, showing that binding strength and insulation can be uncoupled. Single-molecule maps show that nucleosome phasing regularity predicts insulation strength more reliably than CTCF peak intensity. Thus, dif- ferences in motif architecture and TF abundance between species and cell states provide a mechanism for cell-type-specific regulation of CTCF-mediated chromatin insulation.
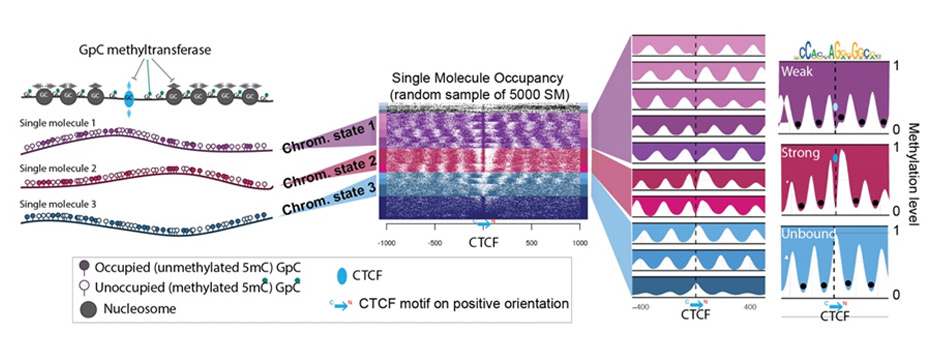 Phasing single-molecule nano-NOMe-seq reveals chromatin state heterogeneity in the context of transcription and long-range interactions
Phasing single-molecule nano-NOMe-seq reveals chromatin state heterogeneity in the context of transcription and long-range interactions
Stephanie Bellini, Catherine Do, Jane A Skok.
A central challenge in molecular biology is determining how 3D chromatin architecture, particularly enhancer-promoter looping and insulating CTCF-mediated interactions, influences gene transcription in individual cells, which has significant implications for healthy and diseased states. To overcome current limitations in imaging and genomic technologies, we developed a cluster-based phasing strategy using long read nano-NOMe-seq to link distinct CTCF binding states—captured at the single molecule level—to the transcriptional status of genes. By stitching partially overlapping long reads and clustering them by shared GpC-accessibility patterns, we stratify CTCF into graded binding states on individual molecules, classify RNA polymerase states at promoters/gene bodies, and infer when spatially separated loci are coordinately activated and occupy loop-competent configurations on the same molecules. When applied to Sox2, Hoxa, and Klf1 regions, cluster-based nano-NOMe-seq phasing reveals how specific topologies bias polymerase behavior and multi-locus activity in ways that bulk assays or locus-engineered imaging cannot fully capture.
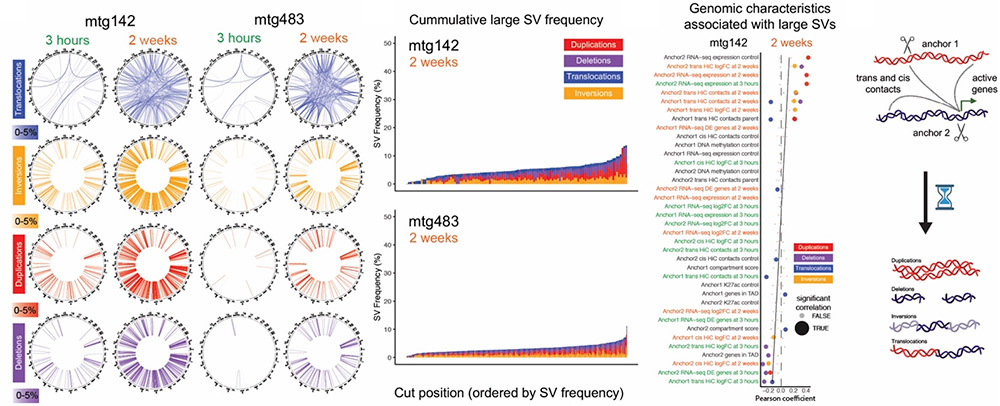 Persistent chromatin alterations and gene expression reprogramming follow widespread DNA damage in glioblastoma
Persistent chromatin alterations and gene expression reprogramming follow widespread DNA damage in glioblastoma
Aram S. Modrek, Ken Chandradoss, Catherine Do, Ravesanker Ezhilarasan, Theodore Sakellaropoulos, Jerome Karp, Yingwen Ding, Ze-Yan Zhang, Melanie Graciani, Giulia Cova, Jennifer E. Phillips-Cremins, Erik P. Sulman, Jane A Skok.
DNA damage from routine cellular processes or exogenous insults can have a lasting impact on gene regulation beyond genetic mutations. The prevailing paradigm for the consequences of DNA damage repair revolves around restoration of the original genetic sequence, but long-term changes in chromatin configuration, gene expression and DNA modifications have not been analyzed. We introduced numerous, simultaneous Cas9-mediated DNA double strand breaks (DSBs) at defined locations in human glioblastoma cells and tracked both non-genetic and genetic alterations over time. Megabase-scale genomic alterations that endured two weeks after the initial damage were detected, involving a shift from transiently increased intra-TAD interactions to persistent long range cis and trans contacts, alterations in gene-expression and associated large structural variations. These findings reveal that widespread DNA damage, such as chemotherapy or radiotherapy, can trigger long-term genetic and non-genetic modifications which alter cellular function and may impact tumor outcome and the emergence of resistant cells.
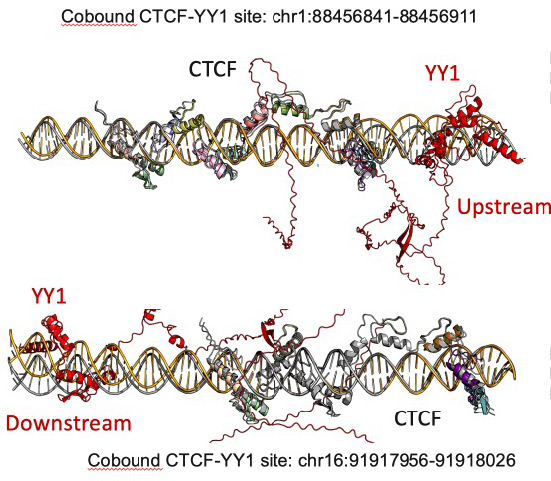 The coordination between CTCF, cohesin and TFs impacts nucleosome repositioning and chromatin insulation to define state specific 3D chromatin folding
The coordination between CTCF, cohesin and TFs impacts nucleosome repositioning and chromatin insulation to define state specific 3D chromatin folding
Catherine Do, Guimei Jiang, Giulia Cova, Paul Zappile, Adriana Heguy, Jane A Skok.
CTCF-mediated chromatin folding plays a key role in gene regulation, however the mechanisms controlling chromatin organization across cell states are not fully elucidated. Comprehensive analyses reveal that CTCF binding stability and cohesin overlap in mice and humans, are regulated by species specific differences in CTCF binding site (CBS) accessibility and enrichment of motifs corresponding to expressed TFs. By analyzing TFs, we confirm the co-operativity and competitiveness of TF/CTCF binding, which we further validate by allele specific analysis of SNPs. TF motif enrichment at CTCF bound sites is determined by cell state-specific transcriptional programs, which either stabilize or destabilize CTCF binding, as reflected by changing TF concentration. To examine CTCF binding in the context of nucleosome positioning, we performed single molecule nano-NOMe-seq. Subsetting reveals a continuum of binding states for CTCF, which are differentially represented at accessible versus inaccessible CBSs. As expected, CTCF degradation leads to a progressive loss of binding and nucleosome repositioning giving profiles similar to CTCF free CBSs. We also observe a similar time dependent effect when the cohesin subcomponent, SSC1 is degraded although CTCF remains bound, indicating that cohesin mediates CTCF-associated nucleosome repositioning. Stratified analysis of CTCF signal strength and accessibility reveals that in the presence of cohesin, CTCF strength contributes to nucleosome repositioning and chromatin insulation independent of accessibility. However, cobound TFs can uncouple the relationship between signal strength and nucleosome repositioning, without affecting the connection between repositioning and insulation. These studies identify mechanisms underlying cell state-specific CTCF profiles, linked to local and long-range chromatin organization.
Factors that determine cell type-specific CTCF binding in health and disease
Catherine Do and Jane A. Skok.
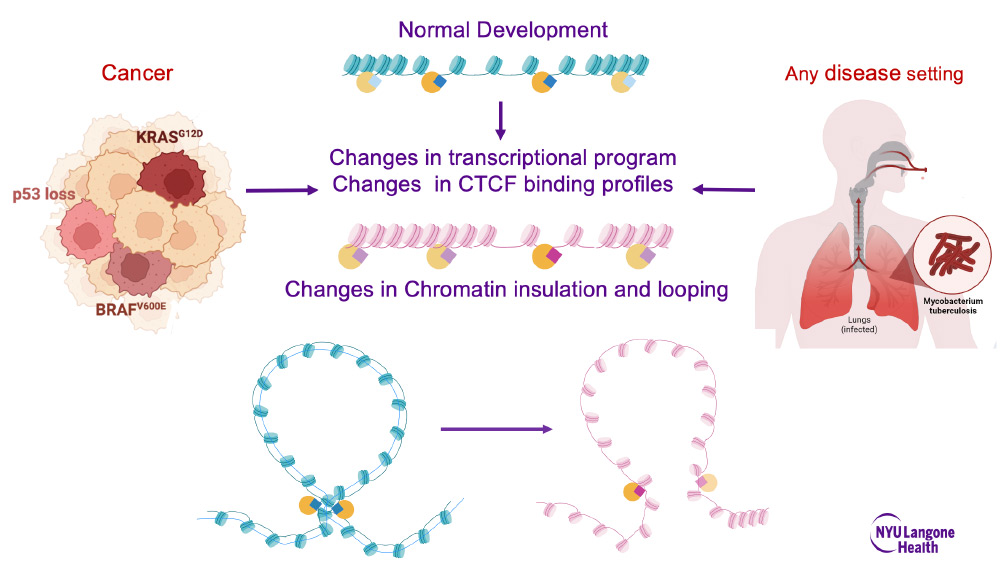
A number of factors contribute to cell type-specific CTCF chromatin binding, but how they act in concert to determine binding stability and functionality has not been fully elucidated. In this review, we tie together different layers of regulation to provide a holistic view of what is known. What emerges from these studies is a multifaceted system in which DNA sequence, DNA and chromatin accessibility, and cell type-specific transcription factors together contribute to CTCF binding profile and function. We discuss these findings in the light of disease settings in which changes in the chromatin landscape and transcriptional programming can disrupt CTCF's binding profile and involvement in looping.
 Binding domain mutations provide insight into CTCF’s relationship with chromatin and its contribution to gene regulation
Binding domain mutations provide insight into CTCF’s relationship with chromatin and its contribution to gene regulation
Catherine Do, Guimei Jiang, Christos C. Katsifis, Domenic N. Narducci, Jie Yang, Giulia Cova, Theodore Sakellaropoulos, Raphael Vidal, Priscillia Lhoumaud, Faye Fara Regis, Nata Kakabadze, Elphege P Nora, Marcus Noyes, Xiaodong Chen, Anders S. Hansen, and Jane A. Skok.
Although only a fraction of CTCF motifs are bound in any cell type, and few occupied sites overlap cohesin, the mechanisms underlying cell-type specific attachment and ability to function as a chromatin organizer remain unknown. To investigate the relationship between CTCF and chromatin we applied a combination of imaging, structural and molecular approaches, using a series of brain and cancer associated CTCF mutations that act as CTCF perturbations. We demonstrate that binding and the functional impact of WT and mutant CTCF depend not only on the unique binding properties of each protein, but also on the genomic context of bound sites and enrichment of motifs for expressed TFs abutting these sites. Our studies also highlight the reciprocal relationship between CTCF and chromatin, demonstrating that the unique binding properties of WT and mutant proteins have a distinct impact on accessibility, TF binding, cohesin overlap, chromatin interactivity and gene expression programs, providing insight into their cancer and brain related effects.
 MethNet: a robust approach to identify regulatory hubs and their distal targets in cancer
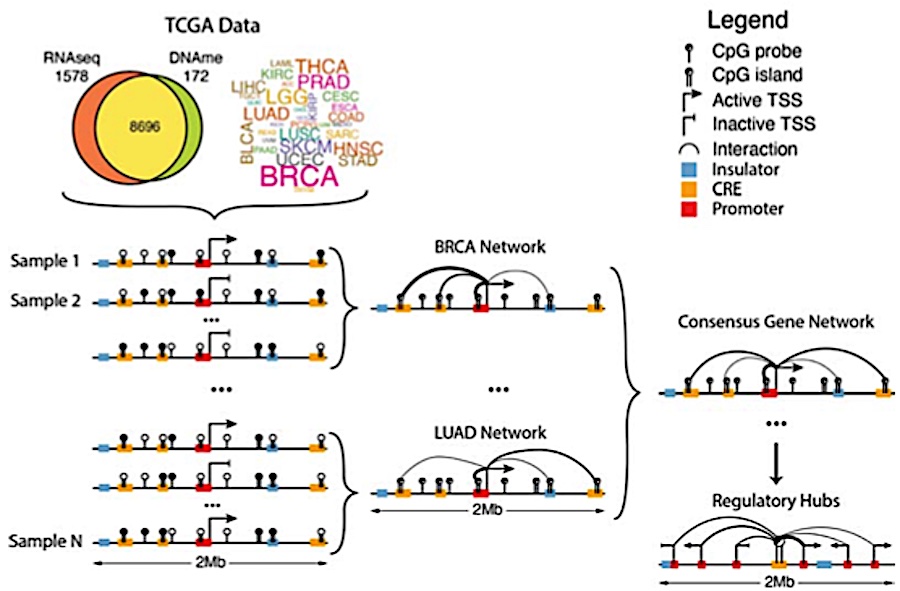
MethNet: a robust approach to identify regulatory hubs and their distal targets in cancer
Theodore Sakellaropoulos, Catherine Do, Guimei Jiang, Giulia Cova, Peter Meyn, Dacia Dimartino, Sitharam Ramaswami, Adriana Heguy, Aristotelis Tsirigos, and Jane A Skok.
Aberrations in the capacity of DNA/chromatin modifiers and transcription factors to bind non-coding regions can lead to changes in gene regulation and impact disease phenotypes. However, identifying distal regulatory elements and connecting them with their target genes remains challenging. Here, we present MethNet, a pipeline that integrates large-scale DNA methylation and gene expression data across multiple cancers, to uncover novel cis regulatory elements (CREs) in a 1Mb region around every promoter in the genome. MethNet identifies clusters of highly ranked CREs, referred to as ‘hubs’, which contribute to the regulation of multiple genes and significantly affect patient survival. Promoter-capture Hi-C confirmed that highly ranked associations involve physical interactions between CREs and their gene targets, and CRISPRi based scRNA Perturb-seq validated the functional impact of CREs. Thus, MethNet-identified CREs represent a valuable resource for unraveling complex mechanisms underlying gene expression, and for prioritizing the verification of predicted non-coding disease hotspots.
 Defining the relative and combined contribution of CTCF and CTCFL to genomic regulation
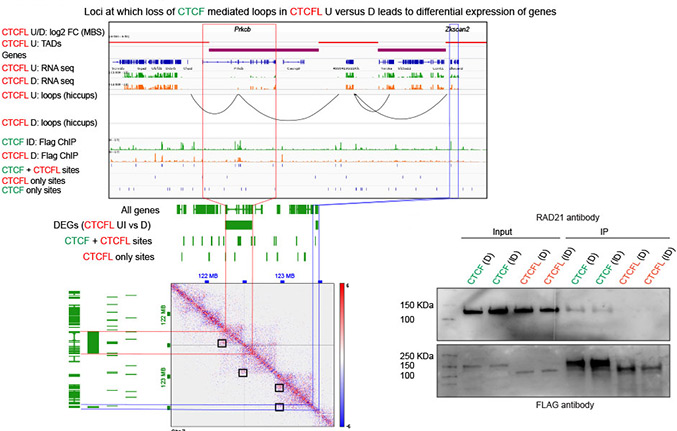
Defining the relative and combined contribution of CTCF and CTCFL to genomic regulation
Mayilaadumveettil Nishana, Caryn Ha, Javier Rodriguez-Hernaez, Ali Ranjbaran, Erica Chio, Elphege P. Nora, Sana B. Badri, Andreas Kloetgen, Benoit G. Bruneau, Aristotelis Tsirigos and Jane A. Skok.
Ubiquitously expressed CTCF is involved in numerous cellular functions, such as organizing chromatin into TAD structures. In contrast, its paralog, CTCFL, is normally only present in the testis. However, it is also aberrantly expressed in many cancers. While it is known that shared and unique zinc finger sequences in CTCF and CTCFL enable CTCFL to bind competitively to a subset of CTCF binding sites as well as its own unique locations, the impact of CTCFL on chromosome organization and gene expression has not been comprehensively analyzed in the context of CTCF function. Using an inducible complementation system, we analyze the impact of expressing CTCFL and CTCF-CTCFL chimeric proteins in the presence or absence of endogenous CTCF to clarify the relative and combined contribution of CTCF and CTCFL to chromosome organization and transcription. We demonstrate that the N terminus of CTCF interacts with cohesin which explains the requirement for convergent CTCF binding sites in loop formation. By analyzing CTCF and CTCFL binding in tandem, we identify phenotypically distinct sites with respect to motifs, targeting to promoter/intronic intergenic regions and chromatin folding. Finally, we reveal that the N, C, and zinc finger terminal domains play unique roles in targeting each paralog to distinct binding sites to regulate transcription, chromatin looping, and insulation. This study clarifies the unique and combined contribution of CTCF and CTCFL to chromosome organization and transcription, with direct implications for understanding how their co-expression deregulates transcription in cancer.
 NSD2 overexpression drives clustered chromatin and transcriptional changes in a subset of insulated domains
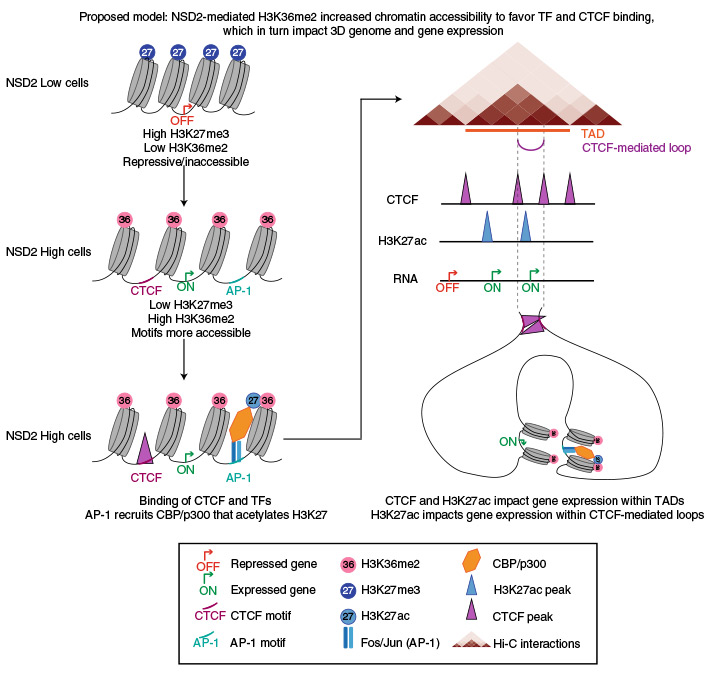
NSD2 overexpression drives clustered chromatin and transcriptional changes in a subset of insulated domains
Priscillia Lhoumaud,Sana Badri,Javier Rodriguez-Hernaez,Theodore Sakellaropoulos,Gunjan Sethia,Andreas Kloetgen,MacIntosh Cornwell,Sourya Bhattacharyya,Ferhat Ay,Richard Bonneau,Aristotelis Tsirigos & Jane A. Skok
CTCF and cohesin play a key role in organizing chromatin into topologically associating domain (TAD) structures. Disruption of a single CTCF binding site is sufficient to change chromosomal interactions leading to alterations in chromatin modifications and gene regulation. However, the extent to which alterations in chromatin modifications can disrupt 3D chromosome organization leading to transcriptional changes is unknown. In multiple myeloma, a 4;14 translocation induces overexpression of the histone methyltransferase, NSD2, resulting in expansion of H3K36me2 and shrinkage of antagonistic H3K27me3 domains. Using isogenic cell lines producing high and low levels of NSD2, here we find oncogene activation is linked to alterations in H3K27ac and CTCF within H3K36me2 enriched chromatin. A logistic regression model reveals that differentially expressed genes are significantly enriched within the same insulated domain as altered H3K27ac and CTCF peaks. These results identify a bidirectional relationship between 2D chromatin and 3D genome organization in gene regulation.
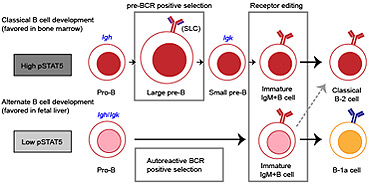 B-1a cells acquire their unique characteristics by bypassing the pre-BCR selection stage
B-1a cells acquire their unique characteristics by bypassing the pre-BCR selection stage
Jason B. Wong, Susannah L. Hewitt, Lynn M. Heltemes-Harris, Malay Mandal, Kristen Johnson, Klaus Rajewsky, Sergei B. Koralov, Marcus R. Clark, Michael A. Farrar & Jane A. Skok
B-1a cells are long-lived, self-renewing innate-like B cells that predominantly inhabit the peritoneal and pleural cavities. In contrast to conventional B-2 cells, B-1a cells have a receptor repertoire that is biased towards bacterial and self-antigens, promoting a rapid response to infection and clearing of apoptotic cells. Although B-1a cells are known to primarily originate from fetal tissues, the mechanisms by which they arise has been a topic of debate for many years. Here we show that in the fetal liver versus bone marrow environment, reduced IL-7R/STAT5 levels promote immunoglobulin kappa gene recombination at the early pro-B cell stage. As a result, differentiating B cells can directly generate a mature B cell receptor (BCR) and bypass the requirement for a pre-BCR and pairing with surrogate light chain. This ‘alternate pathway’ of development enables the production of B cells with self-reactive, skewed specificity receptors that are peculiar to the B-1a compartment. Together our findings connect seemingly opposing lineage and selection models of B-1a cell development and explain how these cells acquire their unique properties.
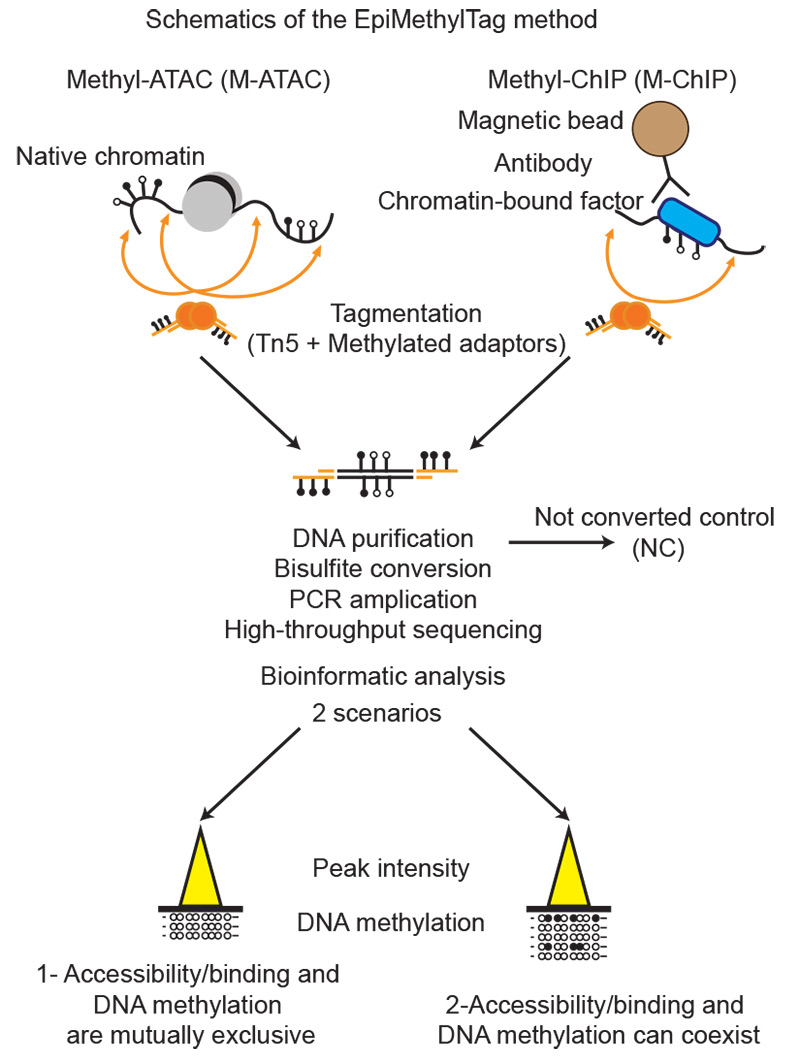 EpiMethylTag: simultaneous detection of ATAC-seq or ChIP-seq signals with DNA methylation
EpiMethylTag: simultaneous detection of ATAC-seq or ChIP-seq signals with DNA methylation
Priscillia Lhoumaud, Gunjan Sethia, Franco Izzo, Theodore Sakellaropoulos, Valentina Snetkova, Simon Vidal, Sana Badri, Macintosh Cornwell, Dafne Campigli Di Giammartino, Kyu-Tae Kim, Effie Apostolou, Matthias Stadtfeld, Dan Avi Landau & Jane Skok
Activation of regulatory elements is thought to be inversely correlated with DNA methylation levels. However, it is difficult to determine whether DNA methylation is compatible with chromatin accessibility or transcription factor (TF) binding if assays are performed separately. We developed a fast, low-input, low sequencing depth method, EpiMethylTag, that combines ATAC-seq or ChIP-seq (M-ATAC or M-ChIP) with bisulfite conversion, to simultaneously examine accessibility/TF binding and methylation on the same DNA. Here we demonstrate that EpiMethylTag can be used to study the functional interplay between chromatin accessibility and TF binding (CTCF and KLF4) at methylated sites.
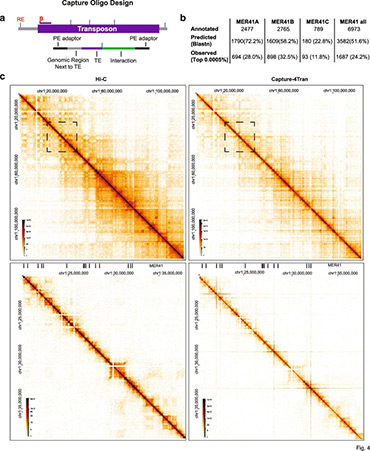 Analysis of 3D genomic interactions identifies candidate host genes that transposable elements potentially regulate
Analysis of 3D genomic interactions identifies candidate host genes that transposable elements potentially regulate
Ramya Raviram, Pedro P. Rocha, Vincent M. Luo, Emily Swanzey,Emily R. Miraldi, Edward B. Chuong, Cédric Feschotte, Richard Bonneau, and Jane A. Skok
The organization of chromatin in the nucleus plays an essential role in gene regulation. About half of the mammalian genome comprises transposable elements. Given their repetitive nature, reads associated with these elements are generally discarded or randomly distributed among elements of the same type in genome-wide analyses. Thus, it is challenging to identify the activities and properties of individual transposons. As a result, we only have a partial understanding of how transposons contribute to chromatin folding and how they impact gene regulation.
Using PCR and Capture-based chromosome conformation capture (3C) approaches, collectively called 4Tran, we take advantage of the repetitive nature of transposons to capture interactions from multiple copies of endogenous retrovirus (ERVs) in the human and mouse genomes. With 4Tran-PCR, reads are selectively mapped to unique regions in the genome. This enables the identification of transposable element interaction profiles for individual ERV families and integration events specific to particular genomes. With this approach, we demonstrate that transposons engage in long-range intra-chromosomal interactions guided by the separation of chromosomes into A and B compartments as well as topologically associated domains (TADs). In contrast to 4Tran-PCR, Capture-4Tran can uniquely identify both ends of an interaction that involve retroviral repeat sequences, providing a powerful tool for uncovering the individual transposable element insertions that interact with and potentially regulate target genes.
4Tran provides new insight into the manner in which transposons contribute to chromosome architecture and identifies target genes that transposable elements can potentially control.
Bibliography
- Stephanie Bellini, Catherine Do & Jane A Skok. (2025). Phasing single-molecule nano-NOMe-seq reveals chromatin state heterogeneity in the context of transcription and long-range interactions. BioRxiv.
- Catherine Do, G. J., Paul Zappile, Adriana Heguy, Jane A. Skok. The coordination between CTCF, cohesin and TFs impacts nucleosome repositioning and chromatin insulation to define state specific 3D chromatin folding. BioRxiv (2025).
- K. C. Aram S. Modrek, Catherine Do, Ravesanker Ezhilarasan, Theodore Sakellaropoulos, Jerome Karp, Yingwen Ding, Ze-Yan Zhang, Melanie Graciani, Giulia Cova, Jennifer E. Phillips-Cremins, Erik P. Sulman and Jane A. Skok. Persistent chromatin alterations and gene expression reprogramming follow widespread DNA damage in glioblastoma. BioRxiv (2025).
- Do C, Jiang G, Cova G, Katsifis CC, Narducci DN, Sakellaropoulos T, Vidal R, Lhoumaud P, Tsirigos A, Regis FFD, Kakabadze N, Nora EP, Noyes M, Hansen AS, Skok JA. Binding domain mutations provide insight into CTCF’s relationship with chromatin and its ability to act as a chromatin organizer. Biorxiv. 2024. Epub 20240925. doi: 10.1101/2024.01.11.575070. PubMed PMID: 38370764; PMCID: PMC10871189.
- Catarina Sacristan; Ben A Youngblood; Peiyuan Lu; Alexander P.R. Bally; Xiaojin (Jane) Xu; Katelyn McGary; Susannah L Hewitt; Jeremy M Boss; Jane A Skok; Rafi Ahmed; Michael L DustinChronic viral infection alters PD-1 locus subnuclear localization in cytotoxic CD8+ T cells. Cell Reports, 2024.
- Theodore Sakellaropoulos Guimei Jiang, Giulia Cova, Peter Meyn, Dacia Dimartino, Sitharam Ramaswami, Adriana Heguy, Aristotelis Tsirigos, Catherine Do, Jane A Skok. MethNet: a robust approach to identify regulatory hubs and their distal targets in cancer. Nature Communications, 2024.
- Tan J, Shenker-Tauris N, Rodriguez-Hernaez J, Wang E, Sakellaropoulos T, Boccalatte F, Thandapani P, Skok J, Aifantis I, Fenyo D, Xia B, Tsirigos A. Cell-type-specific prediction of 3D chromatin organization enables high-throughput in silico genetic screening. Nature biotechnology. 2023. Epub 2023/01/10. doi: 10.1038/s41587-022-01612-8. PubMed PMID: 36624151.
- Li, J; Lhoumaud, P; Riva, A; Piper, C; Dupere-Richer, D; Casellas, Roman H; Sobh, A; Kaestner, C L; Swaroop, A; Kulis, M; Bennett, R L; Skok, J; Licht, J D. 'Dysregulation of Epigenetic Landscape Uncovered the Mechanisms Underlying the Relapse of Pediatric Acute Lymphoblastic Leukemia with NSD2 Mutation'. Blood. 2021 Conference:(63rd):3297- (# 5099022) [Meeting Abstract]
- Ortabozkoyun H, Huang PY, Cho H, Narendra V, LeRoy G, Gonzalez-Buendia E, Skok JA, Tsirigos A, Mazzoni EO, Reinberg D. CRISPR and biochemical screens identify MAZ as a cofactor in CTCF-mediated insulation at Hox clusters. Nat Genet. 2022;54(2):202-12. Epub 2022/02/12. doi: 10.1038/s41588-021-01008-5. PubMed PMID: 35145304.
- Dr. Chewei Anderson Chang , Dr. Jayu Jen , Mr. Shaowen Jiang , Mr. Azin Sayad , Dr. Arvind S. Mer , Dr. Kevin R. Brown , Dr. Allison M. Nixon , Ms. Avantika Dhabaria , Dr. Kwan Ho Tang , Dr. David Venet , Dr. Christos Sotiriou , Dr. Jiehui Deng , Dr. Kwok-Kin Wong , Sylvia Adams , Mr. Peter Meyn , Dr. Adriana Heguy , Dr. Jane A. Skok , Dr. Aristotelis Tsirigos , Dr. Beatrix Ueberheide , Dr. Jason Moffat , Dr. Abhyudai Singh , Dr. Benjamin Haibe-Kains , Mr. Alireza Khodadadi-Jamayran. Ontogeny and Vulnerabilities of Drug-Tolerant Persisters in HER2+ Breast Cancer. Cancer Discovery. (2021)
- Lhoumaud P, Skok J. Simultaneous Tagmentation-Based Detection of ChIP/ATAC Signal with Bisulfite Sequencing. Methods Mol Biol. 2021;2351:337-52. Epub 2021/08/13. doi: 10.1007/978-1-0716-1597-3_19. PubMed PMID: 34382199.
- Zhou, M., L. Kuruvilla, X. Shi, S. Viviano, I. M. Ahearn, C. R. Amendola, W. Su, S. Badri, J. Mahaffey, N. Fehrenbacher, J. Skok, J. Schlessinger, B. E. Turk, D. A. Calderwood and M. R. Philips (2020). "Scaffold association factor B (SAFB) is required for expression of prenyltransferases and RAS membrane association." Proc Natl Acad Sci U S A.
- Vidal, S. E., A. Polyzos, K. Chatterjee, L. S. Ee, E. Swanzey, J. Morales-Valencia, H. Wang, C. N. Parikh, B. Amlani, S. Tu, Y. Gong, V. Snetkova, J. A. Skok, A. Tsirigos, S. Kim, E. Apostolou and M. Stadtfeld (2020). "Context-Dependent Requirement of Euchromatic Histone Methyltransferase Activity during Reprogramming to Pluripotency." Stem Cell Reports.
- Vasudevaraja, V., J. H. Rodriguez, C. Pelorosso, K. Zhu, A. M. Buccoliero, M. Onozato, H. Mohamed, J. Serrano, L. Tredwin, M. Garonzi, C. Forcato, B. Zeck, S. Ramaswami, J. Stafford, A. Faustin, D. Friedman, E. T. Hidalgo, D. Zagzag, J. Skok, A. Heguy, L. Chiriboga, V. Conti, R. Guerrini, A. J. Iafrate, O. Devinsky, A. Tsirigos, J. G. Golfinos and M. Snuderl (2020). "Somatic Focal Copy Number Gains of Noncoding Regions of Receptor Tyrosine Kinase Genes in Treatment-Resistant Epilepsy." J Neuropathol Exp Neurol.
- Aslam, M. A., M. F. Alemdehy, B. Hao, P. H. L. Krijger, C. E. J. Pritchard, I. de Rink, F. I. Muhaimin, I. Nurzijah, M. van Baalen, R. M. Kerkhoven, P. C. M. van den Berk, J. A. Skok and H. Jacobs (2020). "The Ig heavy chain protein but not its message controls early B cell development." Proc Natl Acad Sci U S A 117(49): 31343-31352.
- Mayilaadumveettil Nishana, Caryn Ha, Javier Rodriguez-Hernaez, Ali Ranjbaran, Erica Chio, Elphege P. Nora, Sana B. Badri, Andreas Kloetgen, Benoit G. Bruneau, Aristotelis Tsirigos and Jane A. Skok. ¬Defining the relative and combined contribution of CTCF and CTCFL to genomic regulation. Genome Biology, 21, 108, 2020.
- Fragliasso, V., Verma, A., Manzotti, G., Tameni, A., Bareja, R., Heavican, T. B., Iqbal, J., Wang, R., Fiore, D., Mularoni, V., Chan, W. C., Lhoumaud, P., Skok, J., Zanetti, E., Merli, F., Ciarrocchi, A., Elemento, O. and Inghirami, G.The novel lncRNA BlackMamba controls the neoplastic phenotype of ALK(-) anaplastic large cell lymphoma by regulating the DNA helicase HELLS. Leukemia, 2020.
- Izzo F, Lee SC, Poran A, Chaligne R, Gaiti F, Gross B, Murali R, Deochand S, Ang C, Jones PW, Nam AS, Kim KT, Kothen-Hill S, Schulman RC, Ki M, Lhoumaud P, Skok JA, Viny AD, Levine RL, Kenigsberg E, Abdel-Wahab O, Landau DA. DNA methylation disruption reshapes the hematopoietic differentiation landscape. Nature Genetics, 2020.
- Wong, J. B., S. L. Hewitt, L. M. Heltemes-Harris, M. Mandal, K. Johnson, K. Rajewsky, S. B. Koralov, M. R. Clark, M. A. Farrar and J. A. Skok (2019). "B-1a cells acquire their unique characteristics by bypassing the pre-BCR selection stage." Nat Commun 10(1): 4768.
- Lhoumaud, P., G. Sethia, F. Izzo, T. Sakellaropoulos, V. Snetkova, S. Vidal, S. Badri, M. Cornwell, D. C. Di Giammartino, K. T. Kim, E. Apostolou, M. Stadtfeld, D. A. Landau and J. Skok (2019). "EpiMethylTag: simultaneous detection of ATAC-seq or ChIP-seq signals with DNA methylation." Genome Biol 20(1): 248.
- Lhoumaud, P., S. Badri, J. Rodriguez-Hernaez, T. Sakellaropoulos, G. Sethia, A. Kloetgen, M. Cornwell, S. Bhattacharyya, F. Ay, R. Bonneau, A. Tsirigos and J. A. Skok (2019). "NSD2 overexpression drives clustered chromatin and transcriptional changes in a subset of insulated domains." Nat Commun 10(1): 4843.
- Cantor, D. J., King, B., Blumenberg, L., DiMauro, T., Aifantis, I., Koralov, S. B., Skok, J. A. and David, G. (2019). "Impaired Expression of Rearranged Immunoglobulin Genes and Premature p53 Activation Block B Cell Development in BMI1 Null Mice." Cell Rep 26(1): 108-118 e104.
- Saldana-Meyer, R., Rodriguez-Hernaez, J., Escobar, T., Nishana, M., Jacome-Lopez, K., Nora, E. P., Bruneau, B. G., Tsirigos, A., Furlan-Magaril, M., Skok, J. and Reinberg, D. RNA Interactions Are Essential for CTCF-Mediated Genome Organization Mol Cell, doi:10.1016/j.molcel.2019.08.015 (2019).
- , Rocha, P. P., Luo, V. M., Swanzey, E., Miraldi, E. R., Chuong, E. B., Feschotte, C., Bonneau, R. and Skok, J. A. (2018). "Analysis of 3D genomic interactions identifies candidate host genes that transposable elements potentially regulate." Genome Biol 19(1): 216. PMCID: PMC6292174.
- Oksuz, O., Narendra, V., Lee, C. H., Descostes, N., LeRoy, G., Raviram, R., Blumenberg, L., Karch, K., Rocha, P. P., Garcia, B. A., Skok, J. A. and Reinberg, D. (2018). "Capturing the Onset of PRC2-Mediated Repressive Domain Formation." Mol Cell 70(6): 1149-1162 e1145.
- Issuree, P. D., Day, K., Au, C., Raviram, R., Zappile, P., Skok, J. A., Xue, H. H., Myers, R. M. and Littman, D. R. (2018). "Stage-specific epigenetic regulation of CD4 expression by coordinated enhancer elements during T cell development." Nat Commun 9(1): 3594.
- Cantor, D. J., King, B., Blumenberg, L., DiMauro, T., Aifantis, I., Koralov, S. B., Skok, J. A. and David, G. (2019). "Impaired Expression of Rearranged Immunoglobulin Genes and Premature p53 Activation Block B Cell Development in BMI1 Null Mice." Cell Rep 26(1): 108-118 e104.
- Modrek AS, Golub D, Khan T, Bready D, Prado J, Bowman C, Deng J, Zhang G, Rocha PP, Raviram R, Lazaris C, Stafford JM, LeRoy G, Kader M, Dhaliwal J, Bayin NS, Frenster JD, Serrano J, Chiriboga L, Baitalmal R, Nanjangud G, Chi AS, Golfinos JG, Wang J, Karajannis MA, Bonneau RA, Reinberg D, Tsirigos A, Zagzag D, Snuderl M, Skok JA, Neubert TA, Placantonakis DG. Low-Grade Astrocytoma Mutations in IDH1, P53, and ATRX Cooperate to Block Differentiation of Human Neural Stem Cells via Repression of SOX2. Cell Rep. 2017 Oct 31;21(5):1267-1280. doi: 10.1016/j.celrep.2017.10.009. PubMed PMID: 29091765.
- Hewitt SL, Wong JB, Lee JH, Nishana M, Chen H, Coussens M, Arnal SM, Blumenberg LM, Roth DB, Paull TT, Skok JA. The Conserved ATM Kinase RAG2-S365 Phosphorylation Site Limits Cleavage Events in Individual Cells Independent of Any Repair Defect. Cell reports. 2017;21(4):979-93. Epub 2017/10/27. doi: 10.1016/j.celrep.2017.09.084. PubMed PMID: 29069605; PMCID: PMC5662208.
- Campos-Sanchez, E. et al. Wolf-Hirschhorn Syndrome Candidate 1 Is Necessary for Correct Hematopoietic and B Cell Development. Cell reports 19, 1586-1601, doi:10.1016/j.celrep.2017.04.069 (2017).
- Le Noir, S. et al. The IgH locus 3' cis-regulatory super-enhancer co-opts AID for allelic transvection. Oncotarget, doi:10.18632/oncotarget.14585 (2017).
- Aranda-Orgilles, B., Saldana-Meyer, R., Wang, E., Trompouki, E., Fassl, A., Lau, S., Mullenders, J., Rocha, P.P., Raviram, R., Guillamot, M., Sanchez-Diaz, M., Wang, K., Kayembe, C., Zhang, N., Amoasii, L., Choudhuri, A., Skok, J.A., Schober, M., Reinberg, D., Sicinski, P., Schrewe, H., Tsirigos, A., Zon, L.I. & Aifantis, I. MED12 Regulates HSC-Specific Enhancers Independently of Mediator Kinase Activity to Control Hematopoiesis. Cell stem cell (2016).
- Jiang, T., Raviram, R., Snetkova, V., Rocha, P.P., Proudhon, C., Badri, S., Bonneau, R., Skok, J.A*. & Kluger, Y*. Identification of multi-loci hubs from 4C-seq demonstrates the functional importance of simultaneous interactions. Nucleic Acids Res (2016). * Joint last authors
- Pedro P. Rocha, Ramya Raviram, Yi Fu, JungHyun Kim, Vincent Luo, Arafat Aljoufi, Emily Swanzey, Alessandra Pasquarella, Alessia Balestrini, Emily R. Miraldi, Richard Bonneau, John Petrini, Gunnar Schotta and Jane A. Skok. A damage independent role for 53BP1 that impacts break order and Igh architecture during CSR. Cell reports. doi:10.1016/j.celrep.2016.05.073 (2016). PMID: 27320916.
- Charlotte Proudhon, Valentina Snetkova, Ramya Raviram, Camille Lobry, Sana Badri, Tingting Jiang, Bingtao Hao, Thomas Trimarchi, Yuval Kluger, Iannis Aifantis, Richard Bonneau, Jane A Skok. Active and inactive enhancers co-operate to exert localized and long-range control of gene regulation. Cell reports, (2016). PMCID: PMC4899175.
- Yi Fu, Pedro P. Rocha, Vincent M. Luo, Ramya Raviram, Yan Deng, Esteban O. Mazzon & Jane A. Skok. CRISPR-dCas9 and sgRNA scaffolds enable dual-colour live imaging of satellite sequences and repeat-enriched individual loci. Nature communications 7 (2016).
- Thomas-Claudepierre, A.S., Robert, I., Rocha, P.P., Raviram, R., Schiavo, E., Heyer, V., Bonneau, R., Luo, V.M., Reddy, J.K., Borggrefe, T., Skok, J.A. & Reina-San-Martin, B. Mediator facilitates transcriptional activation and dynamic long-range contacts at the IgH locus during class switch recombination. J Exp Med (2016).
- Raviram, R., Rocha, P.P., M¸ller, C.L., Miraldi, E.R., Badri, S., Fu, Y., Swanzey, E., Proudhon, C., Snetkova, V., Bonneau, R. & Skok, J.A. 4C-ker: A Method to Reproducibly Identify Genome-Wide Interactions Captured by 4C-Seq Experiments. PLoS computational biology 12, e1004780 (2016).
- Sun, A., Novobrantseva, T.I., Coffre, M., Hewitt, S.L., Jensen, K., Skok, J.A., Rajewsky, K., and Koralov, S.B. (2015). VH replacement in primary immunoglobulin repertoire diversification. Proc Natl Acad Sci U S A 112, E458-466.
- Mullenders, J., Aranda-Orgilles, B., Lhoumaud, P., Keller, M., Pae, J., Wang, K., Kayembe, C., Rocha, P.P., Raviram, R., Gong, Y., et al. (2015). Cohesin loss alters adult hematopoietic stem cell homeostasis, leading to myeloproliferative neoplasms. J Exp Med.
- Narendra V, Rocha PP, An D, Raviram R, Skok JA, et al. Transcription. CTCF establishes discrete functional chromatin domains at the Hox clusters during differentiation. Science. 2015 Feb 27;347(6225):1017-21.
- Sun, A., Novobrantseva, T., Coffre, M., Hewitt, S.L., Jensen, K., Skok, J., Rajewsky, K. & Koralov, S., : VH replacement in primary immunoglobulin repertoire diversification. Proc Natl Acad Sci U S A 112, E458-466 (2015).
- Dose, Marei; Emmanuel, Akinola Olumide; Chaumeil, Julie; Zhang, Jiangwen; Sun, Tianjiao; Germar, Kristine; Aghajani, Katayoun; Davis, Elizabeth M; Keerthivasan, Shilpa; Bredemeyer, Andrea L; Sleckman, Barry P; Rosen, Steven T; Skok, Jane A; Le Beau, Michelle M; Georgopoulos, Katia; Gounari, Fotini. 'beta-Catenin induces T-cell transformation by promoting genomic instability'. Proceedings of the National Academy of Sciences of the United States of America. 2014 111(1):391-396 (# 781772).
- Rocha, P.P., Micsinai, M., Kluger, Y. & Skok, J.A. Response to Casellas et al. The Origin of Recurrent Chromosomal Translocations: Different approaches for analyzing genome wide data sets can lead to differences in results. Mol Cell 51, 277-278 (2013).
- Chaumeil, J., Micsinai, M., Ntziachristos, P., Roth, D.B., Aifantis, I., Kluger, Y., Deriano, L. & Skok, J.A. The RAG2 C-terminus and ATM protect genome integrity by controlling antigen receptor gene cleavage. Nat Commun 4, 2231 (2013).
- Julie Chaumeil, Mariann Micsinai, Panagiotis Ntziachristos, Ludovic Deriano, Joy M-H Wang, Yanhong Ji, Elphege P. Nora, Matthew J. Rodesch, Jeffrey A. Jeddeloh, Iannis Aifantis, Yuval Kluger, David G. Schatz, & Jane A. Skok. Higher-order looping and nuclear organization of antigen receptor loci facilitate targeted RAG cleavage and regulated rearrangement in recombination centers. Cell Reports 3 (2) 359-70 (2013). (Highlighted in Cell Leading Edge, Cell 152, March14, 2013.)
- Rocha, P.P., Micsinai, M., Kim, J.R., Hewitt, S.L., Souza, P.P., Trimarchi, T., Strino, F., Parisi, F., Kluger, Y. & Skok, J.A. Close Proximity to Igh Is a Contributing Factor to AID-Mediated Translocations. Molecular Cell (2012).
- Johnson K, Chaumeil J, Micsinai M, Wang JM, Ramsey LB, Baracho GV, Rickert RC, Strino F, Kluger Y, Farrar MA and Skok, J.A. IL-7 Functionally Segregates the Pro-B Cell Stage by Regulating Transcription of Recombination Mediators across Cell Cycle. J.Immunol., doi:10.4049/jimmunol.1200368 (2012). (Featured “In This Issue” section of The Journal of Immunology).
- Yougui Xiang, Xiaorong Zhou, Susannah L. Hewitt, Jane A. Skok, and William T. Garrard. (2011). A multifunctional element in the mouse Ig locus that specifies repertoire and Ig loci subnuclear location. J. Immunol.
- Collins, A., Hewitt, S.L., Chaumeil, J., Sellars, M., Micsinai, M., Allinne, J., Parisi, F., Nora, E.P., Bolland, D.J., Corcoran, A.E., Kluger, Y., Bosselut, R., Ellmeier, W., Chong, M.M., Littman, D.R. & Skok, J.A. RUNX Transcription Factor-Mediated Association of Cd4 and Cd8 Enables Coordinate Gene Regulation. Immunity 34, 303-314 (2011).
- Deriano, L., Chaumeil, J., Coussens, M., Multani, A., Chou, Y., Alekseyenko, A.V., Chang, S., Skok, J.A. & Roth, D.B. The RAG2 C terminus suppresses genomic instability and lymphomagenesis. Nature 471, 119-123 (2011).
- Susannah L. Hewitt, Bu Yin, Yanhong Ji, Julie Chaumeil, Katarzyna Marszalek, Jeannette Tenthorey, Georgia Salvagiotto, Natalie Steinel, Laura Ramsey, Jacques Glysdael, Michael Farrar, Barry P. Sleckman, David G. Schatz, Meinrad Busslinger, Craig H. Bassing and Jane A. Skok. RAG1 and ATM co-ordinate mono-allelic recombination and nuclear positioning of immunoglobulin loci. Nat Immunol. 2009 Jun;10(6):655-64. (Highlighted in Cell Leading Edge, Cell 137, June 12, 2009 p. 981.) PMID: 19448632
- Karnowski A, Cao C, Matthias G, Carotta S, Corcoran LM, Martensson IL, Skok JA, Matthias P. (2008) Silencing and nuclear repositioning of the lambda5 gene locus at the pre-B cell stage requires Aiolos and OBF-1. PLoS ONE 3 (10), e3568.
- Hewitt SL, Farmer D, Marszalek K, Cadera E, Liang HE, Xu Y, Schlissel MS, Skok JA. (2008). Association between the Igk and Igh immunoglobulin loci mediated by the 3' Igk enhancer induces 'decontraction' of the Igh locus in pre-B cells. Nature Immunology Apr;9(4):396-404
- Johnson K, Hashimshony T, Sawai CM, Pongubala JM, Skok JA, Aifantis I, Singh H. (2008). Regulation of Immunoglobulin Light-Chain Recombination by the Transcription Factor IRF-4 and the Attenuation of Interleukin-7 Signaling. Immunity. Mar;28(3):335-45.
- Fitzsimmons SP, Bernstein RM, Max EE, Skok JA, Shapiro MA. Dynamics of Accessibility, Nuclear Positioning, Recombination and Transcription at the Kappa Locus during B-Cell Development. J Immunology 179, 5264-73 (2007).
- Liu H, Schmidt-Supprian M, Shi Y, Hobeika E, Barteneva N, Jumaa H, Pelanda R, Reth M, Skok J, Rajewsky K, Shi Y (2007) Yin Yang 1 is a critical regulator of B-cell development. Genes & Dev 2007, 21:1179-1189.
- Skok JA, Gisler R, Novatchkova M, Farmer D, de Laat W, Busslinger M: Reversible contraction by looping of the Tcra and Tcrb loci in rearranging thymocytes. Nature Immunology 2007, 8:378-387.
- Parker MJ, Licence S, Erlandsson L, Galler GR, Chakalova L, Osborne CS, Morgan G, Fraser P, Jumaa H, Winkler TH, Skok J, Mårtensson IL. (2005) The pre-B-cell receptor induces silencing of VpreB and l5 transcription. EMBO Journal. Nov 16;24(22):3895-905.
- Goldmit M*, Ji Y*, Skok J*, Roldán E, Jung S, Cedar H, Bergman Y. (2005) Epigenetic ontogeny of the k locus during B cell development. Nature Immmunology 6(2):198-203. (*Joint first authorship).
- Roldán E, Fuxa M, Chong W, Martinez D, Novatchkova M, Busslinger M, Skok JA. (2005) Locus decontraction and centromeric recruitment contribute to allelic exclusion of the immunoglobulin heavy-chain gene. Nature Immmunology 6(1):31-41.
- Fuxa M*, Skok J*, Souabni A, Salvagiotto G, Roldan E, Busslinger M. (2004) Pax5 induces V-to-DJ rearrangements and locus contraction of the immunoglobulin heavy-chain gene. Genes & Dev. Feb 15;18(4):411-22. (*Joint first authorship.)
- Thomas RM, Schmedt C, Novelli M, Choi BK, Skok J, Tarakhovsky A, Roes J. (2004) C-terminal SRC kinase controls acute inflammation and granulocyte adhesion. Immunity. Feb;20(2):181-91.
- Kosak ST, Skok JA, Medina KL, Riblet R, Le Beau MM, Fisher AG, Singh H. (2002) Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science. Apr 5;296(5565):158-62.
- Skok JA, Brown KE, Azuara V, Caparros ML, Baxter J, Takacs K, Dillon N, Gray D, Perry RP, Merkenschlager M, Fisher AG. (2001) Nonequivalent nuclear location of immunoglobulin alleles in B lymphocytes. Nature Immunology Sep;2(9):848-54.
- Siepmann K, Skok J, van Essen D, Harnett M, Gray D. (2001) Rewiring of CD40 is necessary for delivery of rescue signals to B cells in germinal centres and subsequent entry into the memory pool. Immunology. Mar;102(3):263-72.
- Skok J, Poudrier J, Gray D. (1999) Dendritic cell-derived IL-12 promotes B cell induction of Th2 differentiation: a feedback regulation of Th1 development. J. Immunology Oct 15;163(8):4284-91.
- Skok J, Solomon E, Reid KB, Thompson RA. (1981) Distinct genes for fibroblast and serum C1q. Nature. Aug 6;292(5823):549-51.

